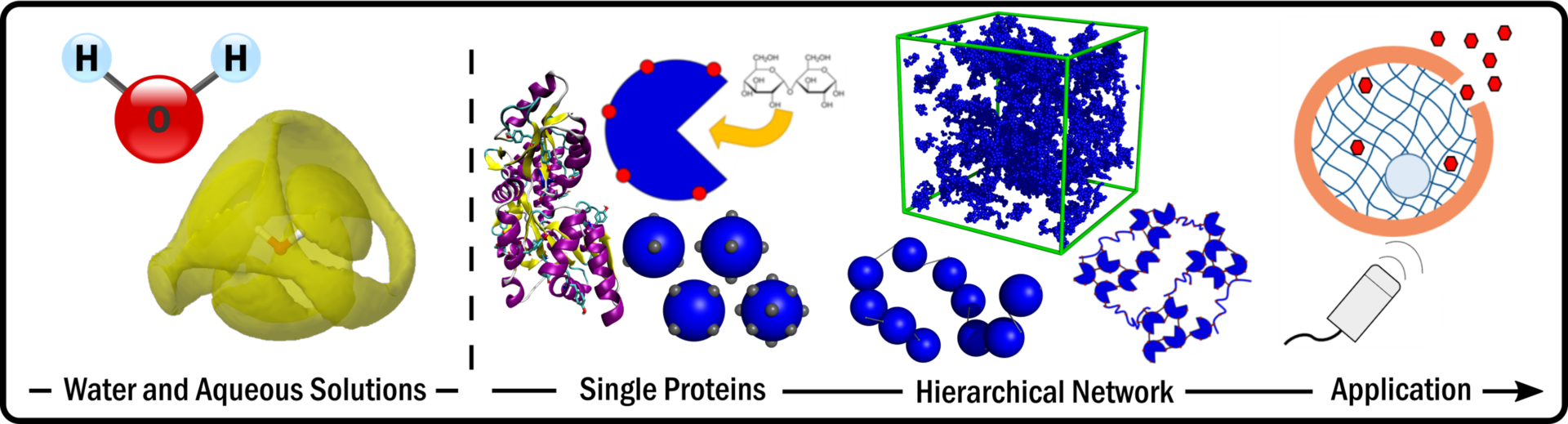
Research
Water structure and dynamics
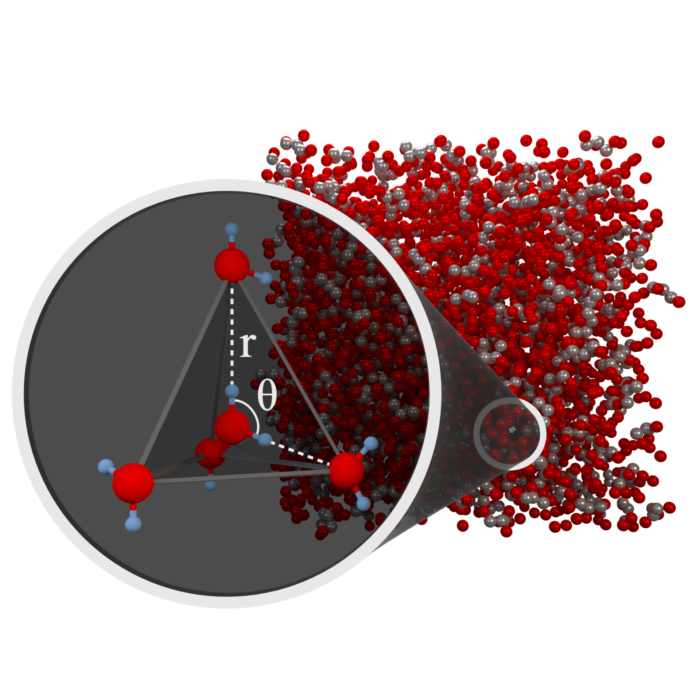
Low density water in a nano-segregated cryoprotectant solution
Water is essential for life. We are interested in the structure and dynamics of water under extreme environmental conditions. Our investigations combine neutron diffraction experiments at the ISIS neutron & muon Facility and computational modelling to determine the atomistic level structure of the liquid mixtures. In collaboration with Prof Mike Ries & Dr Dan Baker at Leeds we explore the dynamics of the same liquids using NMR. Join us for a Virtual weekly Water journal club to learn more.
You can read more about our work on:
- Solute Specific Perturbations to Water Structure and Dynamics in Tertiary Aqueous Solution (HERE)
- TMAO resists the compression of water structure by magnesium perchlorate: terrestrial kosmotrope vs. Martian chaotrope (HERE)
- Temperature-Dependent Segregation in Alcohol–Water Binary Mixtures Is Driven by Water Clustering (HERE)
- Highly compressed water structure observed in a perchlorate aqueous solution (HERE)
and in Science highlight articles
- Frozen frog gives clues to improving freeze storage techniques HERE
- Water surprise: how two molecules can have the same, but also the opposite, effect HERE
- SANDALS diffractometer sheds light on the enigma of Martian channel formation (HERE)
Single molecule manipulation

Single molecule force spectroscopy as a tool to measure mechanical properties
Single molecule force spectroscopy is a powerful tool to explore the response of biological molecules to the application of an applied force. This approach provides valuable insight into biomolecule mechanical stability, as well as details of the underlying energy landscape. We mechanically manipulate proteins from extremophilic organisms to understand how they can remain folded and functional in environmentally extreme conditions. Extremophile proteins provide fascinating model systems for understanding the physics of life.
You can read about our research on
- Differential effects of hydrophobic core packing residues for thermodynamic and mechanical stability of a hyperthermophilic protein (HERE)
- The hierarchical emergence of worm-like chain behaviour from globular domain polymer chains (HERE)
- The physics of pulling polyproteins: a review of single molecule force spectroscopy using the AFM to study protein unfolding (HERE)
Hierarchical biomechanics
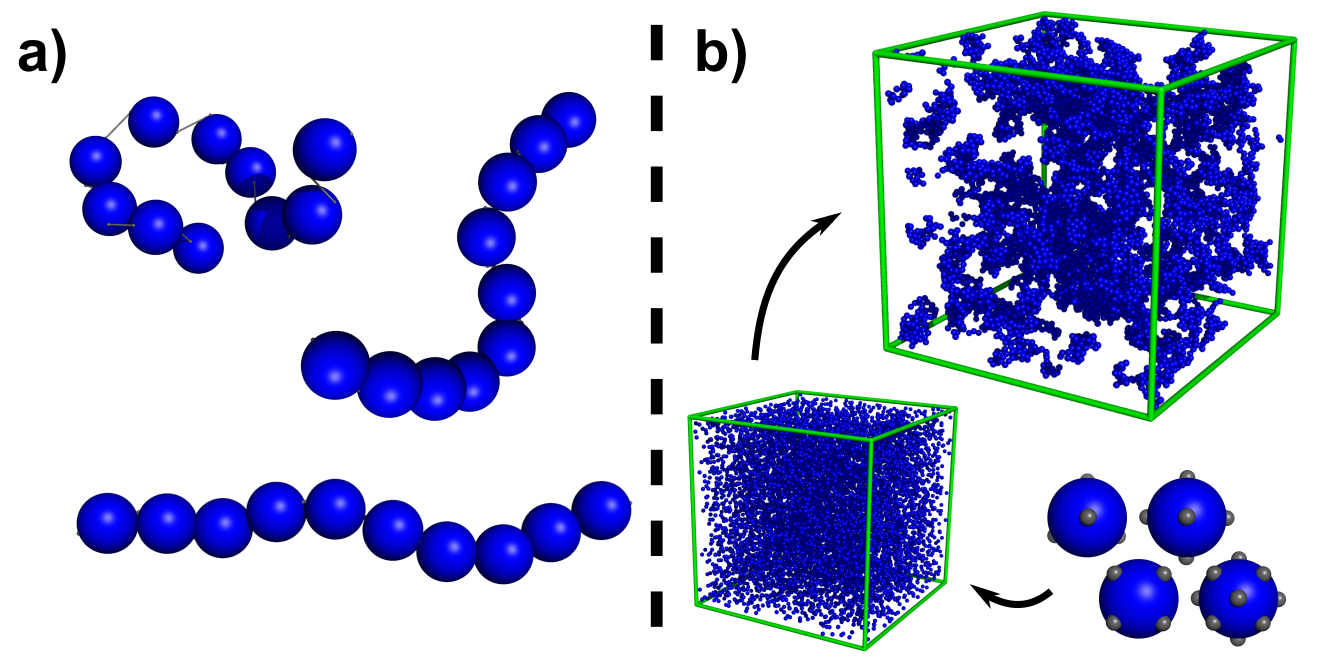

Exploiting Engineered Polyproteins in the Modular Design of Robust, Tuneable and Biofunctional Hydrogels
We are creating folded protein hydrogels which possess specific biological function capabilities, enabling dynamic changes in mechanical and structural properties in response to biomolecular cues. This is achieved through a cross length scale, physics-based approach which will translate knowledge of the nanoscale biophysics of folded proteins to the mesoscale architecture and function of novel folded polyprotein hydrogels. By understanding the physics of the building block (the folded protein) and its connectivity (the polyprotein network) we are creating a platform for the production of novel, biomaterials. You can read about our latest work in this article HERE and in our recent publications which study:
- Control of Nanoscale in situ Protein Unfolding Defines Network Architecture and Mechanics of Protein Hydrogels (HERE)
- Reaction Rate Governs the Viscoelasticity and Nanostructure of Folded Protein Hydrogels (HERE)
- Network Growth and Structural Characteristics of Globular Protein Hydrogels (HERE)
- Single molecule protein stabilisation translates to macromolecular mechanics of protein network (HERE)
- The hierarchical emergence of worm-like chain behaviour from globular domain polymer chains (HERE)
Multi-composite biomaterials
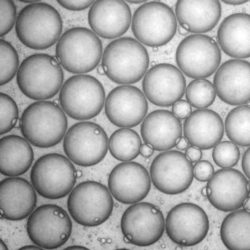
Controlling drug release in targeted drug delivery
Prolonged exposure of chemotherapeutic agents to tumour cells over several cell cycles has been found to be more effective for the treatment of cancer than a short burst of chemotherapeutic agents. We are investigating a composite system involving liposomes with encapsulated nano-hydrogel embedded with nanobubblestermed ‘lipogel’, which will provide a drug transport mechanism with an ultrasound triggered release system. This system mimics the ability of natural cells to take advantage of properties from both the lipid bilayer and nano-hydrogel to create a multifunctional system. The lipid bilayer aims to protect the hydrogel from degradation in the bloodstream, as well as reducing the side effects of toxic drugs on healthy cells by targeting to the region of interest. The presence of a drug loaded gel within the liposome core aims to further control the speed that the drug molecules are released.

